I have an extensive background in freshwater ecology, the dynamics of lower trophic (microbial and zooplankton populations), and molecular techniques for studying lake ecosystems from my graduate studies and post-doctoral work. This leads me to my research focus, which is the gene-to-ecosystem approach to understanding freshwater ecological processes. I aim to bring my diverse perspectives and research expertise to the development of a research program that allows partnerships with national and international organizations and to play a role in assisting Indigenous communities around the world in managing their territories in the face of climate change and anthropogenic impacts.
iTrackDNA: STANDARDIZING TARGETED ENVIRONMENTAL DNA (eDNA) RESOURCES FOR ENVIRONMENTAL MONITORING
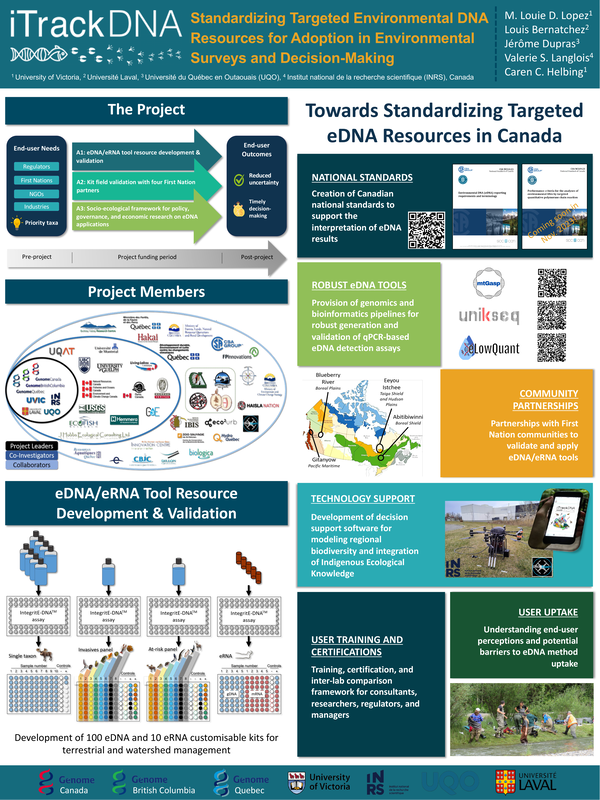
Environmental DNA (eDNA), or genetic material that organisms release into the environment, can provide rapid, non-destructive, accurate and cost-effective biodiversity information. But inconsistent methods and poor eDNA detection tools make it hard for end users (regulators, industry, Indigenous Peoples, NGOs) to use them because they give too many false negatives and false positives, which can make it hard for managers to make good decisions. Here, we provide the latest updates on the iTrackDNA project, a multi-year, large-scale applied research initiative supported by Genome Canada, Genome British Columbia, and Genome Québec that aims to standardize eDNA technology based on targeted quantitative PCR (qPCR) to address issues with researchers and end-users across Canada and industries. This presentation will outline recent efforts designed to increase end-user capacity using cutting-edge, easily accessible, and ethically sound analytical genomics-based targeted eDNA tools for effective decision-making by: 1) supporting the creation of a second national eDNA standard on targeted qPCR-based assay performance criteria; 2) providing annotated mitochondrial genomic resources and new bioinformatic pipelines ('unikseq') to advance tool development; 3) developing validated eDNA and eRNA kits to monitor conservation priority species within Canadian coastal and inland ecosystems; 4) generating decision support software (iTrackDNA mobile app) for regional biodiversity data collection; and 5) constructing eDNA training and inter-lab validation frameworks for consultants, researchers, regulators, and managers. By establishing national eDNA standards and decentralizing specific eDNA resources, iTrackDNA aims to increase the eDNA community of practice and confidently enable eDNA applications in inland and coastal ecological surveys as well as biosurveillance for mining, forestry, energy, and infrastructure projects.
SEDIMENTARY DNA (DNA) IN RECONSTRUCTING FISH SETTLEMENT HISTORY IN ALBERTA'S OIL SANDS REGION
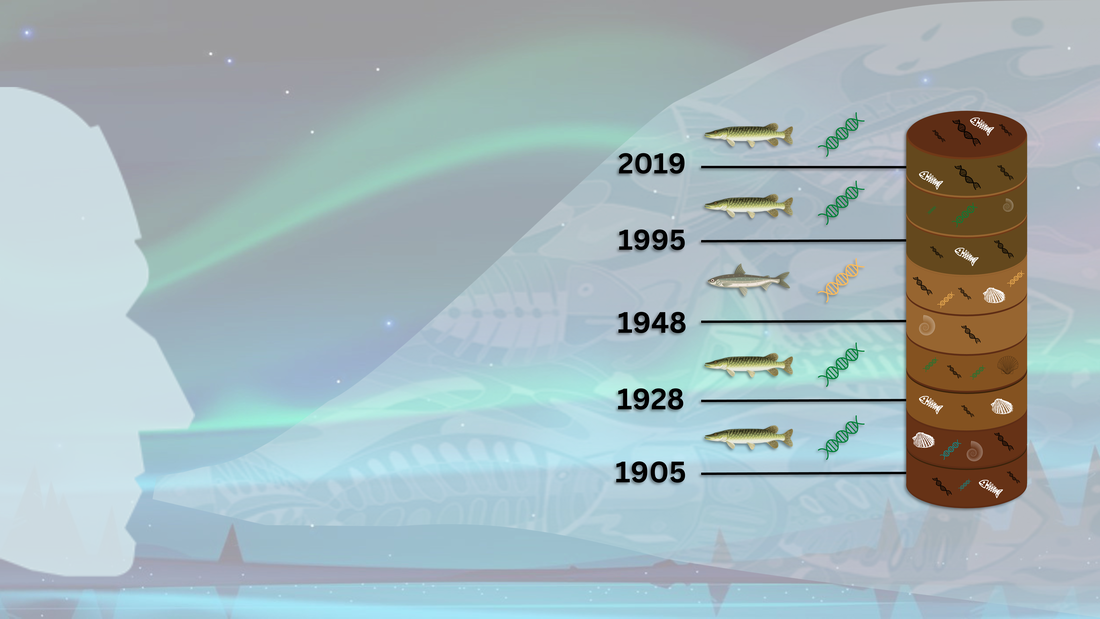

Environmental DNA (eDNA) studies have proliferated over the last decade, with promising data generated to describe the diversity of macro- and micro-organisms in most environments. The recovery of DNA preserved in the sediment of aquatic systems (sedDNA) has provided short-term and long-term data on biological groups (e.g., bacteria, phytoplankton, zooplankton, and fishes) and has advanced our understanding of how environmental changes have affected aquatic communities. Herein, we demonstrate the utility of fish sedDNA from lake sediment cores in reconstructing the paleolimnological history of fish settlement in selected lakes within the Oil Sands region in Alberta. Eight targeted qPCR-based eDNA assays for freshwater fish species were rigorously designed and validated. We also applied a coupled precipitation and column-based approach to effectively isolate and detect sedDNA. With these tools and methods, we found sedDNA from native and non-native freshwater fish species in sediment cores that went back more than a hundred years. This confirmed historical records of introductions by humans. The use of fish sedDNA provided greater temporal resolution into the historical fish faunal records, bridging the knowledge gap that spanned from 100- to 150-year-old data. The present study also provides confirmation of the presence of native fish species in different lakes through sedDNA detection, predating human-mediated introductions in each lake. Our findings enabled the refinement of native freshwater fish ranges and clarification of the effect of human-mediated introductions on fish diversity in aquatic bodies within the Oil Sands region, providing critical baseline information for environmental impact assessments.
ALLOMETRIC SCALING IN COMMUNITY RNA TRANSCRIPTS
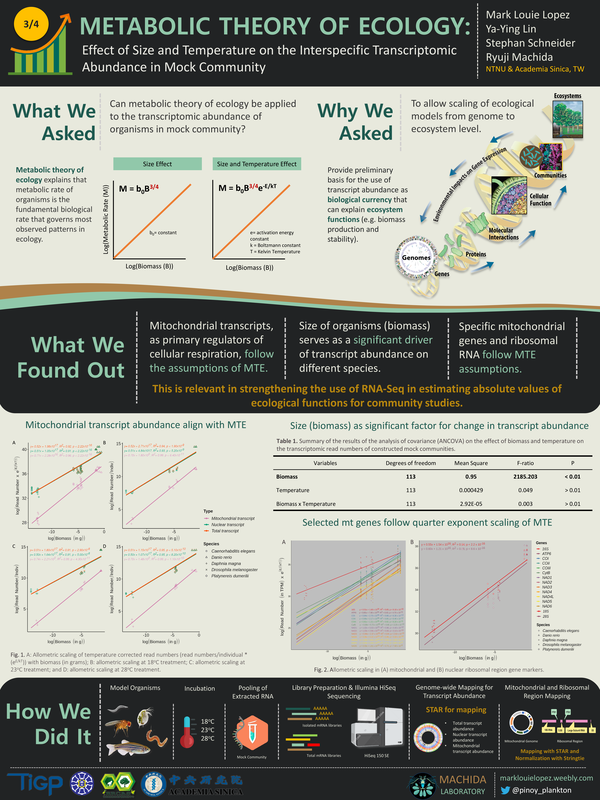
Metatranscriptomics allows the profiling of community messenger RNA (mRNA) and ribosomal RNA (rRNA) transcript abundance under certain environmental conditions. But differences in the amount of RNA transcripts in communities of different sizes are still not fully understood. This makes it harder to use metatranscriptomics in community studies. We looked into the growth-rate hypothesis (GRH) and the metabolic theory of ecology (MTE) to see if the allometric scaling of interspecific RNA transcript (mRNA and rRNA) abundance was true. We did this by doing metatranscriptomic analysis of fake communities made up of model organisms. Results suggest that body size imposes significant constraints on RNA transcript abundance. Interestingly, the relationship between the total mitochondrial transcript abundance (mRNA and rRNA slopes were –0.30 and –0.28, respectively) and body size aligned with the MTE assumptions with slopes close to –¼, while the nuclear transcripts displayed much steeper slopes (mRNA and rRNA slopes were –0.33 and –0.40, respectively). The assumed temperature dependence was not observed in this study. At the gene level, the allometric slopes range from 0 to –1. Overall, the above results showed that larger individuals have lesser RNA transcript abundance per tissue mass than smaller ones regardless of temperature. Analyses of field-collected microcrustacean zooplankton samples demonstrated that the correction of size effect, using the allometric exponents derived from the model organism mock community, explains the patterns of interspecific RNA transcripts abundance within metatranscriptome. By combining allometry with metatranscriptomics, RNA transcript reads can be used to learn more about how ecological processes work in communities that are very complex.
USE OF RNA-BASED WORKFLOW IN ZOOPLANKTON MONITORING
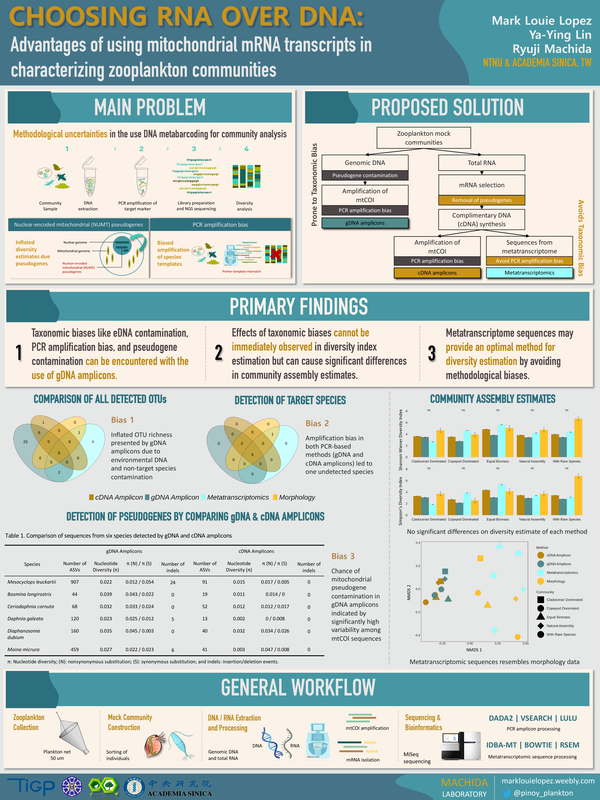

RNA-based methods have been useful in advancing many aspects of plankton research but remain uncommon in zooplankton studies. Methods that capture messenger RNA (mRNA) expression profiles, like metatranscriptomics, allow more thorough monitoring of the temporal changes in zooplankton species diversity and community composition in response to changes in environmental conditions. However, doubts about using the RNA-based approach remain due to misconceptions on the difficulty of sample preservation, storage, and bioinformatics procedures. With this, promoting the advances in RNA-based workflow for zooplankton community studies is needed. Here, we provide best practices in metatranscriptomics workflow, including zooplankton community sample collection and preservation, RNA extraction, RNA quality checking, nucleic acid long-term storage, and bioinformatics analysis. Moreover, we demonstrate the advantages of using an RNA-based approach in avoiding methodological uncertainties (i.e., PCR amplification bias and nuclear-encoded mitochondrial (NUMT) pseudogene contamination) in estimating species richness and composition of zooplankton communities. Results from our study show that metatranscriptomics provides diversity estimates that closely resembled data derived from morphological analysis. Metatranscriptomics has also shown consistency in monitoring temporal changes in the community composition of field-collected samples. Overall, these findings demonstrate the advantages of metatranscriptomics as an effective tool for diversity monitoring in zooplankton research.
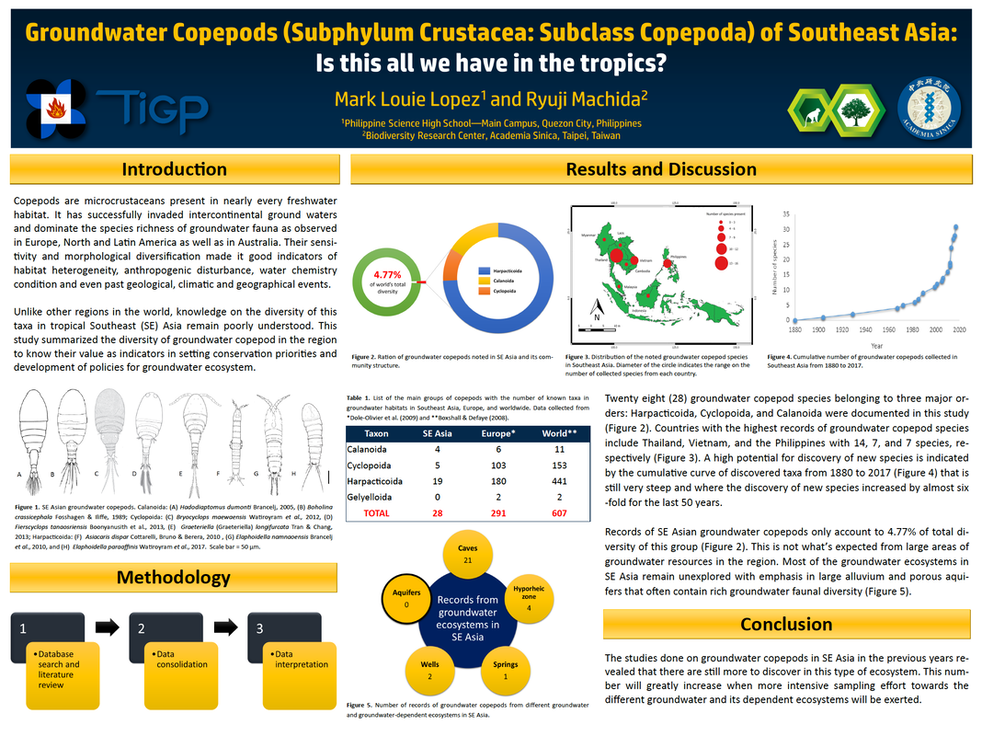
GROUNDWATER MICROCRUSTACEANS IN SOUTHEAST ASIA
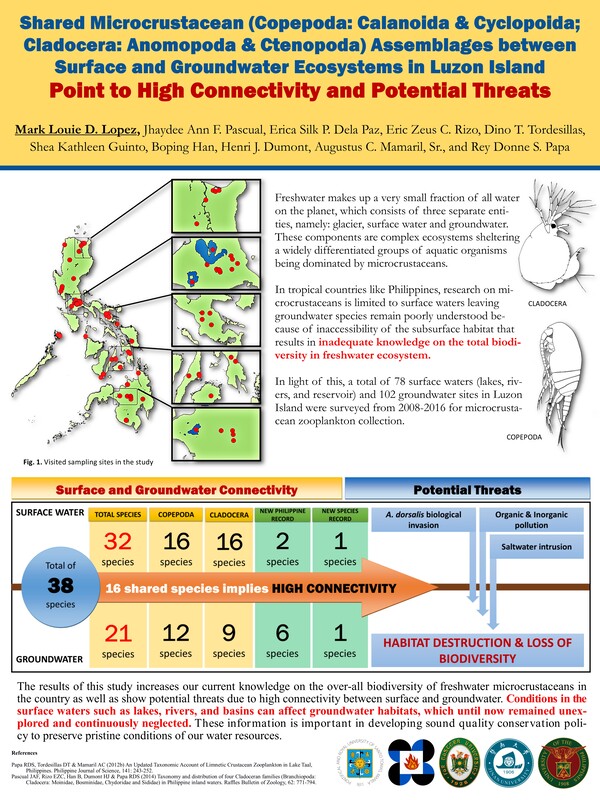
In the Philippines, the ecology of microcrustaceans from groundwaters and their dependent ecosystems remain poorly understood, yet knowledge about this group can elucidate patterns in groundwater biodiversity to develop sound conservation policies. We collect samples of microcrustaceans from groundwater-dependent ecosystems, including artesian wells, caves, springs, and piped groundwater pumps during the dry (Nov 2014 to Apr 2015) and wet (May–Oct 2015) seasons. So far, a total of 21 species from Cladocera and Copepoda, including 2 obligate stygobionts and 19 species consist with surface water and facultative stygobiotic taxa. Significant differences in microcrustacean assemblages were noted among types of groundwater-dependent ecosystems, wherein wells and caves harbored more abundant assemblages with higher total species richness; however, no significant variations were observed between seasons. Furthermore, sampling sites were highly characterized by altitude, specific conductivity, and total water hardness. Microcrustacean assemblages in sampled sites were highly dominated by the influence of temperature, dissolved oxygen, and altitude. Species rarefaction analysis revealed low species richness in sampled sites within the region, supporting the existing notion that temperate groundwater-dependent ecosystems were more diverse, and faunal composition in terms of ecological groups is extremely different in tropical and temperate settings.
In regional scale, our team also tries to document occurrence of groundwater copepod species in Southeast Asia. Copepods have successfully penetrated the groundwater realm through a series of morphological diversifications and adaptations. Research on this taxon has increased over the last decade due to its potential in revealing the status of groundwater environmental health and biodiversity. Despite efforts in documenting this group in other regions, groundwater copepods in Southeast (SE) Asia remain barely studied. To date, only 48 species belonging to 22 genera from Harpacticoida, Cyclopoida, and Calanoida have been documented from groundwater and groundwater-dependent habitats across the region. The steep species accumulation curve from 1980 up to present indicates a high possibility of discovering more new species. Spatial distribution shows high local endemicity than regional scales, where some species considered to be rare and endemic were actually common in local habitats. Overall, the low number of records in the region is due to the lack of experts and limited accessibility to groundwater and dependent ecosystems like aquifers and groundwater wells. A more intensive effort in documenting the diversity and distribution of groundwater copepods and building collaborations between experts in the region is highly needed. This information is important in drafting future conservation and management policies for the groundwater resources in the region.
FRESHWATER ZOOPLANKTON FAUNA OF THE PHILIPPINES
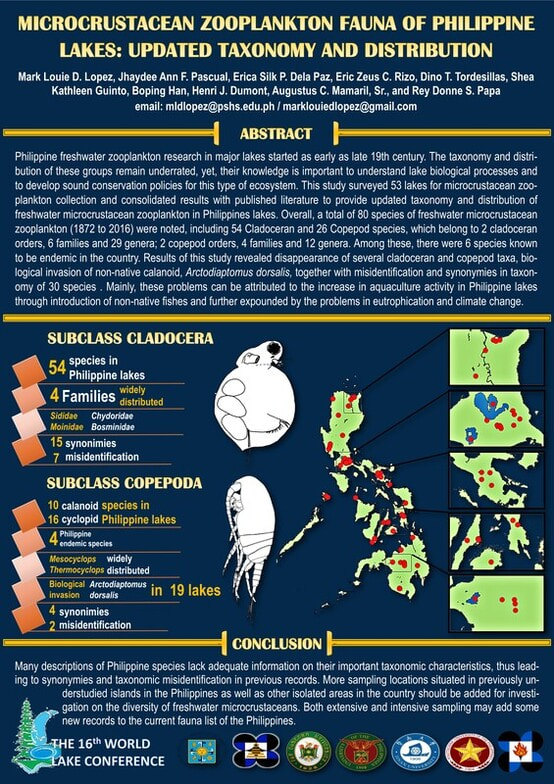
In the past decade, our team under the supervision of Dr. Rey Donne S. Papa of the University of Santo Tomas aims to provide a checklist that puts together available information on the taxonomy, distribution, and current status of freshwater microcrustacean zooplankton in the Philippines. To date, 81 species have been recorded from Philippine inland waters, including 55 cladoceran and 36 copepod species, in two cladoceran orders (six families); and in two copepod orders (four families). The level of endemicity and distribution patterns of microcrustaceans in the archipelago's freshwater systems reflects the island's origins, biogeographical status, and location in the tropics. However, there are problems: in terms of taxonomy, species level identification is often doubtful and further study on systematics and biogeography is needed to settle conflicts in identification. Also, synonymies and misidentifications are also continuously noted in previous Philippine records. In addition, the introduction of non-native species of fishes, zooplankton, and other aquatic organisms has begun negatively impacting inland aquatic biodiversity in the country, which is further exacerbated by eutrophication and other environmental changes.